A non-local diffusion magnetic resonance imaging tract density biomarker to stratify, predict, and interpret survival rates in human glioblastoma
New publication presents a connectomics-based biomarker for predicting glioblastoma survival
A novel connectomics-based biomarker offers a new lens for predicting glioblastoma survival. A new study titled “A non-local diffusion magnetic resonance imaging tract density biomarker to stratify, predict, and interpret survival rates in human glioblastoma” has been published in Neuro-Oncology. The research, co-led by members of the Computational Neuroscience team at Sano and researchers from Humanitas University and IRCCS Humanitas Research Hospital, presents a new approach to predict and understand the clinical outcomes of one of the most lethal brain tumors.
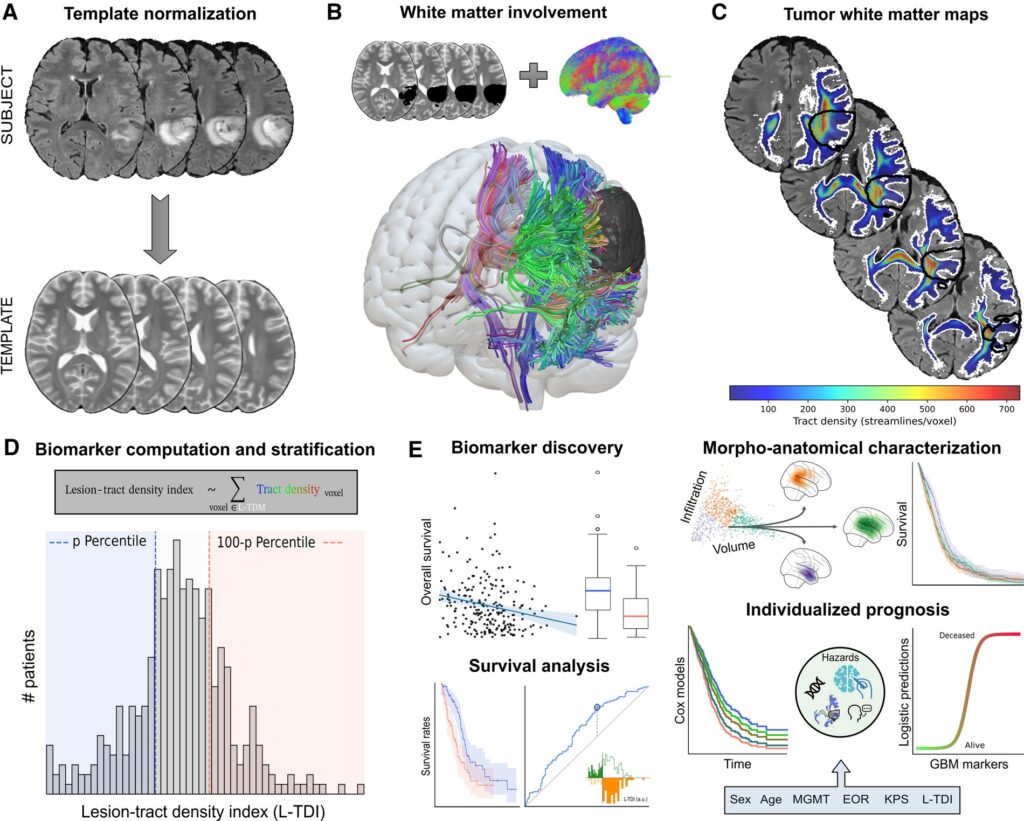
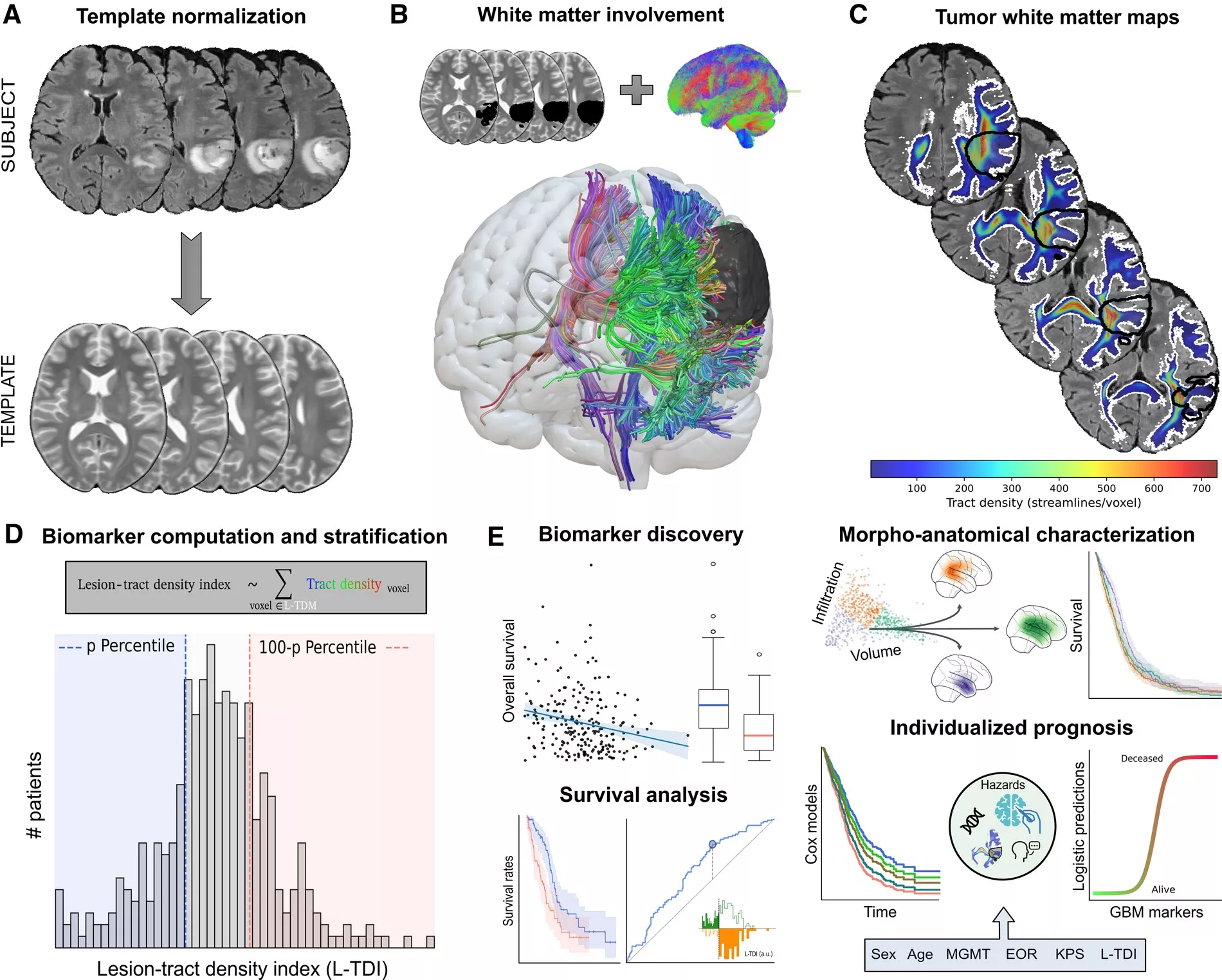
Traditionally, glioblastoma (GBM) has been regarded as a focal lesion disrupting brain tissue. However, increasing evidence suggests that these tumors are active members of brain networks, leveraging white matter as a morphological substrate to grow and migrate to distant regions. This work introduces the Lesion-Tract Density Index (L-TDI), which uses normative white matter tractograms to quantify the intersection between the tumor and the brain’s large-scale network architecture.
By testing the L-TDI in two independent cohorts comprising over 850 patients, the researchers demonstrated that this marker is a robust predictor of survival. Crucially, the L- TDI incorporates information about the tumor’s size, its location, and the density of the white matter it affects, making it more effective than looking at tumor volume alone.
The study found that patients with a high L-TDI, indicating a more complex interaction with the brain’s white matter scaffold, had significantly worse prognoses. This integrative approach provides an anatomically interpretable biomarker that could revolutionize clinical decision-making. For instance, L-TDI maps could help refine radiotherapy planning by identifying potential routes of microscopic tumor spread along white matter pathways.
Ultimately, this work advocates for a paradigm shift toward connectomics-guided neuro- oncology, where treating a brain tumor means understanding its place within the brain’s complex structural network.
Autors: Joan Falcó-Roget (Sano), Gianpaolo Antonio Basile, Anna Janus, Sara Lillo, Letterio S Politi, Jan K. Argasinski, Alberto Cacciola
See the full text: academic.oup.com/neuro-oncology/article/